VOLUME 13 (Supplement)
Philipp. Sci. Lett. 2020 13 (Supplement) 092-101
available online: November 30, 2020
*Corresponding author
Email Address: mpbalolong@up.edu.ph
Date received: March 15, 2020
Date revised: August 26, 2020
Date accepted: October 20, 2020
ARTICLE
Genotyping of β-Lactamase (blaCTX-M, blaOXA, blaSHV, and blaTEM) genes from Extended-spectrum β-Lactamase-producing Enterobacteriaceae Clinical Isolates of Ospital ng Maynila Memorial Medical Center, Philippines
University of the Philippines Manila, Ermita, Manila
1000 Philippines
2Department of Science, College of Arts and Sciences,
Miriam College, Katipunan Avenue, Loyola Heights,
Quezon City 1108 Philippines
3Department of Pathology, Ospital ng Maynila,
Quirino Ave, corner Roxas Blvd., Malate, Manila
1004 Philippines
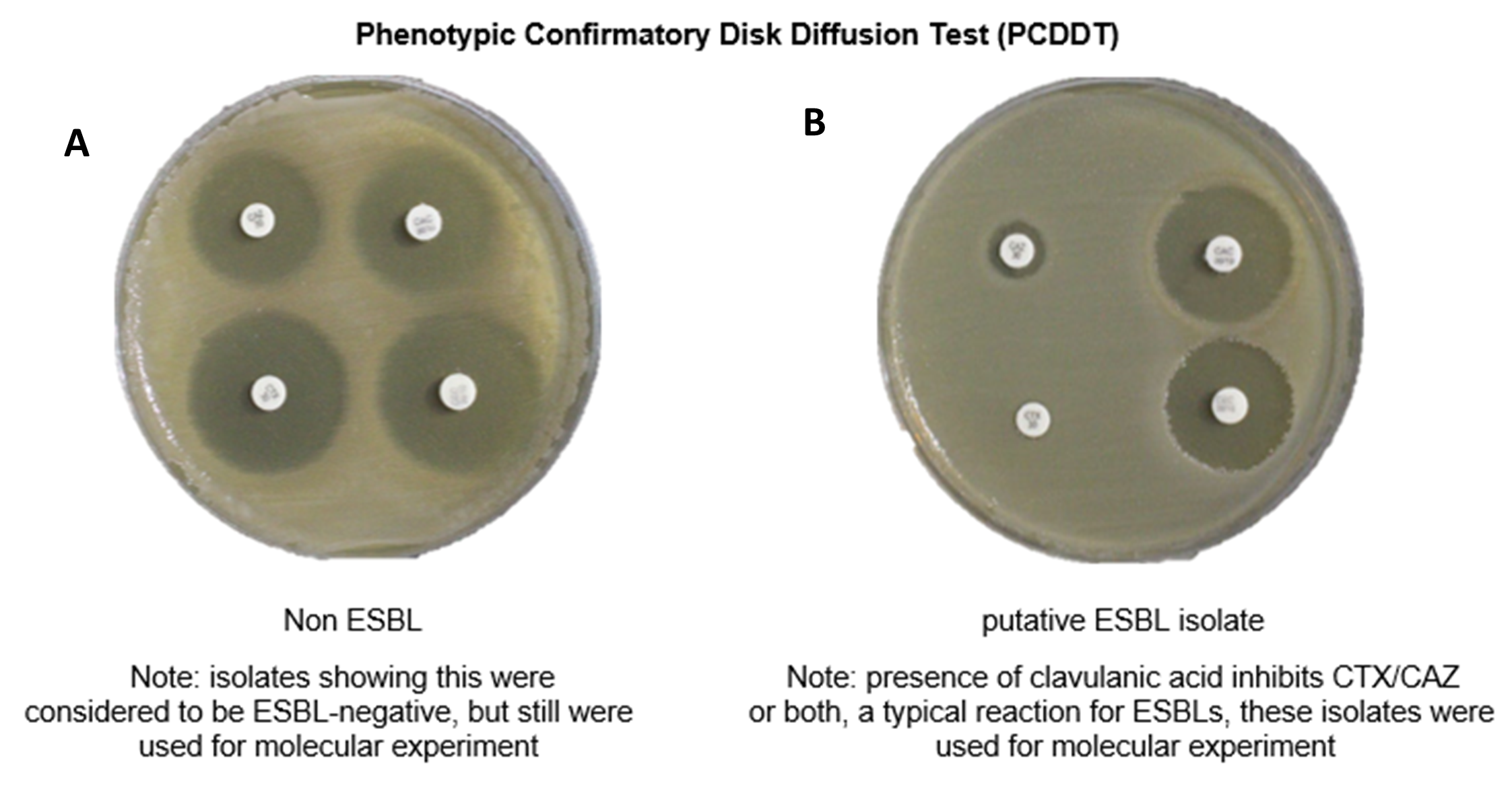
This paper reports the presence of extended-spectrum β-lactamase (ESBL)-producing bacteria exhibiting multiple bla genes from sputum, endotracheal aspartate, urine, corneal specimen, wound discharge, breastmilk, and abscess from infected patients of Ospital ng Maynila Memorial Medical Center. About 55 out of 100 clinical isolates were considered as putative ESBL-producers being resistant to more than one of the β-lactam antibiotics tested. They were further identified as Enterobacter aerogenes, Enterobacter hafnia, Enterobacter spp., Enterobacter cloacae, Klebsiella spp., Klebsiella pneumoniae, Klebsiella rhinoscleromatis, Klebsiella cloacae, Klebsiella ozaenae, and Escherichia coli. Only 40% of the isolates were confirmed for ESBL-production through the Phenotypic Disk Diffusion Test (PCDDT). The type of β-lactamase genes carried by these ESBL-producers was also determined by amplification of the blaCTX-M, blaOXA, blaSHV, and blaTEM genes by Polymerase Chain Reaction. The blaCTX-M gene was the most predominant bla gene followed by the blaOXA gene which was detected in 53/55 (96.36%) of the isolates. On the other hand, 44/55 (80%) of the isolates possessed the blaSHV gene while 38/55 (69.1%) carried the blaTEM gene. The presence of multiple bla genes within the same strain with 35 out of 55 (63.6%) possessing all the four genes tested were also observed. Ten (18.18%) of the strains possessed three of the β-lactamase genes and another 10 harboured at least two genes. Interestingly, bla genes were also detected even for isolates that were negative for the PCDDT indicating that these genes might be present but the β-lactamase enzyme might not have been actively expressed by the bacteria.
© 2026 SciEnggJ
Philippine-American Academy of Science and Engineering