VOLUME 19 NUMBER 1 (January to June 2026)
SciEnggJ. 2026 19 (1) 164-174
available online: 07 April 2026
DOI: https://doi.org/10.54645/2026191PRZ-49
*Corresponding author
Email Address: kizroquel@national-u.edu.ph
Date received: 23 September 2025
Dates revised: 20 November 2025
Date accepted: 29 January 2026
ARTICLE
Molecular identification and phylogenetic analysis of Philippine Fasciola spp. isolates
2Department of Biology, School of Science and Engineering, Ateneo de Manila University, Quezon City 1108 Philippines
3Research Institute for Tropical Medicine, Muntinlupa City 1781 Philippines
4Parasite and Vector Biology and Control Program, University of the Philippines Los Baños Zoonoses Center, Los Baños, Laguna 4031 Philippines
5Office of Research Coordination, University of the East, Manila City 1008 Philippines
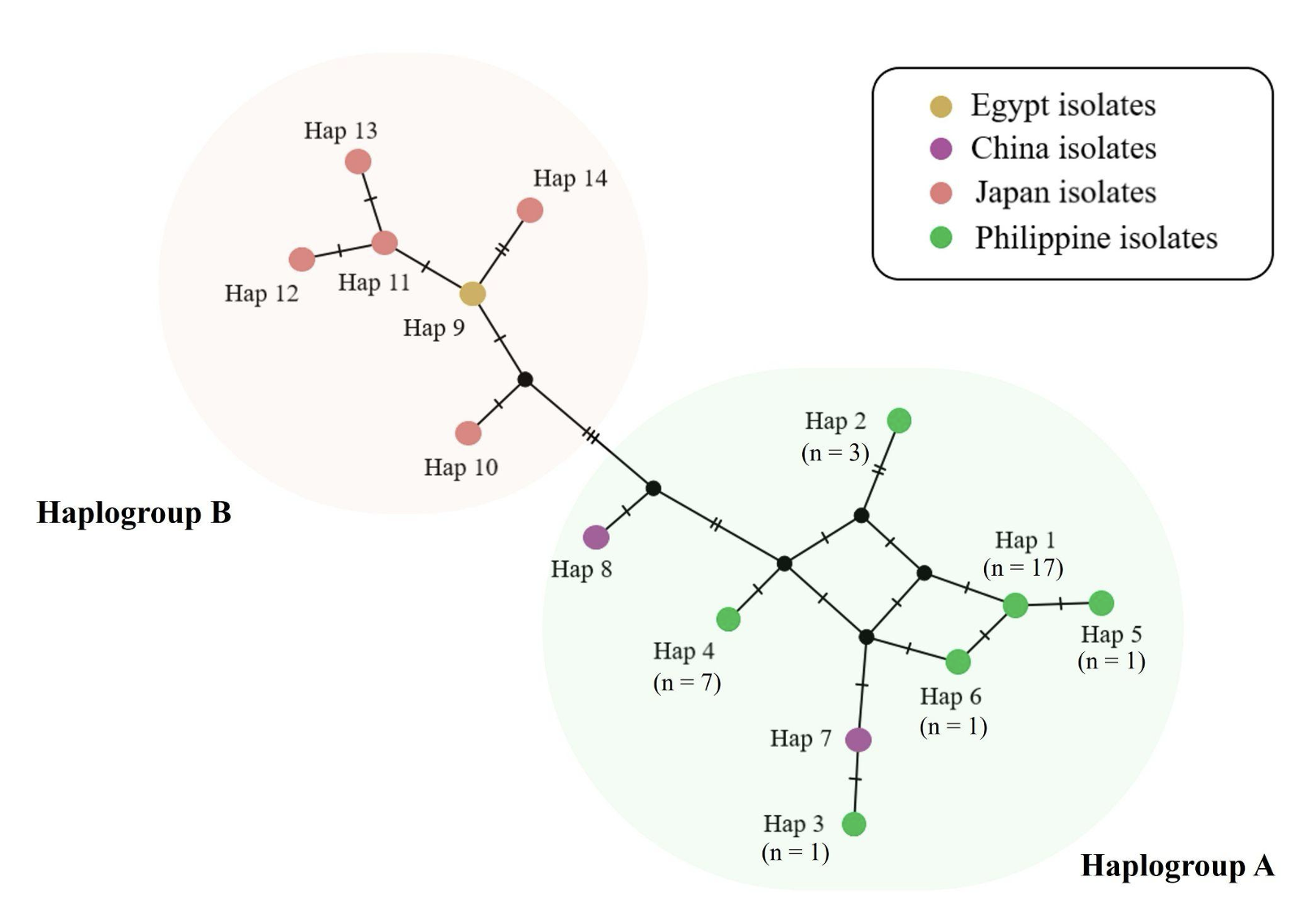
Fascioliasis is a foodborne neglected tropical disease (NTD) caused by trematodes in the genus Fasciola. It is a major cause of morbidity and mortality in cattle and carabaos worldwide. Despite the substantial agricultural impact of bovine fascioliasis in the Philippines, genetic studies on species identification and phylogeny remain limited. This study describes the morphological and genetic characteristics of adult Fasciola sp. collected from swamp buffaloes in the agricultural provinces of Sorsogon and Batangas, Philippines. Morphological analysis was based on body length, cone length, and the presence or absence of shoulders while genetic identification used partial COX1-16S rRNA sequences. Among 33 intact worms, initial morphological assessment on unstained specimens suggested 11 Fasciola gigantica-like (33.3%), 3 Fasciola hepatica-like (9.1%) and 19 Fasciola sp. intermediate forms (57.6%). PCR amplification of a partial COX1-16S rRNA sequence identified F. gigantica in 32 specimens with successful DNA extraction. Phylogenetic analysis of 30 high-quality sequences showed clustering of the Philippine isolates with F. gigantica samples from Basrah, Iraq, and Lanzhou, China. Notably, our analysis is consistent with previous reports suggesting that Philippine swamp buffaloes originated from mainland China. These findings demonstrate the value of molecular approaches for characterizing fasciolid populations and provide supporting evidence on genetic relatedness and possible dispersal patterns.
© 2026 SciEnggJ
Philippine-American Academy of Science and Engineering